 
Recent studies have found that the human microbiome is influenced by various host factors including genotype, lifestyle such as dietary habit, physiological status such as aging, pathophysiological status, and host environment. Accurately identifying and understanding these associations is critical to elucidate the true roles of the microbiome in health and disease states and for development of new diagnostics and therapeutic targets based on the microbiome. The metagenomic sequencing data provide valuable resources for investigating associations between the microbiome and host environmental/clinical factors. This opens a new era of metagenomics studies to explore microbial communities sampled directly from the environments without need for cultivation. The advent of next-generation sequencing (NGS) technology enables the generation of large volume of metagenomic sequencing data at moderate cost. The method has been incorporated into the freely available R package BhGLM ( and ), providing a useful tool for analyzing microbiome data. The results show that the proposed method has desirable properties and outperform the previously used methods in terms of both empirical power and Type I error. We evaluate and demonstrate the proposed method via extensive simulation studies and the application to mouse gut microbiome data.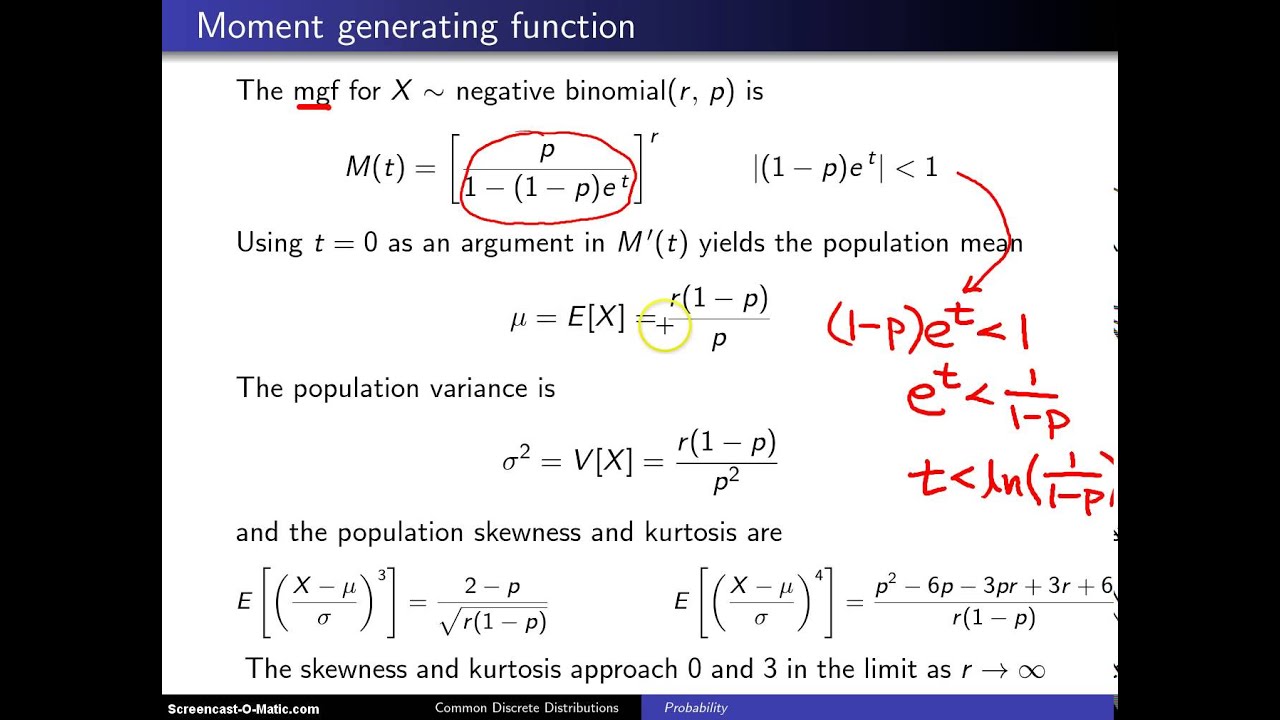  
We have developed a flexible and efficient IWLS (Iterative Weighted Least Squares) algorithm to fit the proposed NBMMs by taking advantage of the standard procedure for fitting the linear mixed models. Although having not dealt with zero-inflation, the proposed mixed-effects models account for correlation among the samples by incorporating random effects into the commonly used fixed-effects negative binomial model, and can efficiently handle over-dispersion and varying total reads. In this article, we propose negative binomial mixed models (NBMMs) for detecting the association between the microbiome and host environmental/clinical factors for correlated microbiome count data. In addition to the well-known properties of microbiome count measurements, for example, varied total sequence reads across samples, over-dispersion and zero-inflation, microbiome studies usually collect samples with hierarchical structures, which introduce correlation among the samples and thus further complicate the analysis and interpretation of microbiome count data. These data provide valuable resources for investigating interactions between the microbiome and host environmental/clinical factors. Recent advances in next-generation sequencing (NGS) technology enable researchers to collect a large volume of metagenomic sequencing data. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
